LC-MS based Targeted Lipidomics of Oxylipins
In the past decade the Schebb lab has developed a comprehensive LC-MS/MS method for the quantification of oxylipins. Together with the ILS oxylipin interest group we developed technical recommendations defining the benchmark for state of the art oxylipin analysis (Schebb et al. 2025). Detailed information how to carry out this analysis and how to address the numerous challenges and pitfalls has been published by our group. However, since the method evolved step by step the information has not been reported in a single article but is distributed over the (supplementary material of) several articles:
A general introduction in LC-MS analysis of oxylipins can be found as part of our reviews (Willenberg et al., 2015; Gladine et al., 2019; Schebb et al., 2022, O´Donnell et al., 2022). The challenges and strategies for analysis of oxylipins are also described in two webinars (youtube-video; video).
Sample preparation
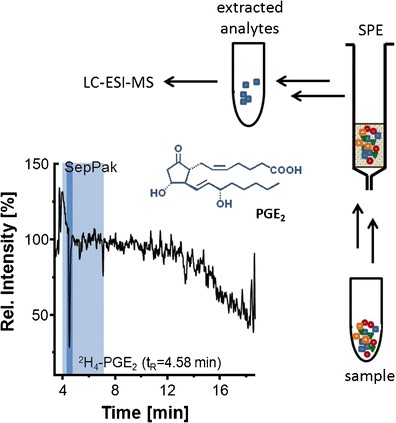
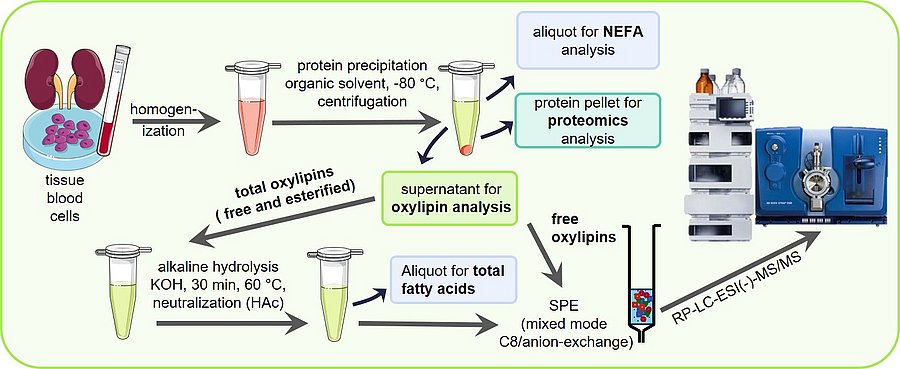
Sample preparation is key for a sound oxylipin analysis. A comparison of different solid phase extraction methods as well as liquid liquid extraction can be found in Ostermann et al., 2015. The optimization of the release of esterified oxylipins by saponification to make them accessible for LC-MS analysis is described in Ostermann et al., 2020. Based on this a standard operation procedure SOP for the analysis of total oxylipins has been published in Koch et al., 2020, excel SI. A more detailed version can be also found in Mainka et al., 2020, excel SI and Rund and Schebb, 2023. An SOP for the analysis of free oxylipins can be found in the article of Kutzner et al., 2019 and again more detailed in Rund and Schebb, 2023.
Using the same sample preparation, free and total fatty acids can be analyzed described by Koch et al., 2021. Different lipid classes, such as free fatty acids, polar lipids and triglycerides, can be separated prior analysis by means of SPE as described by Kutzner et al., 2017.
MS-Detection
The optimization of MS-detection of the oxylipins by electrospray tandem MS is described in detail by Rund et al., 2018 and Kutzner et al., 2019. A list of all oxylipins covered by the method including transitions can be found in these articles and in Hartung et al., 2023. This list can be downloaded here as excel file, together with a method file for a SCIEX 5500 instrument and a SCIEX 6500+ instrument.
Calibration and quantification
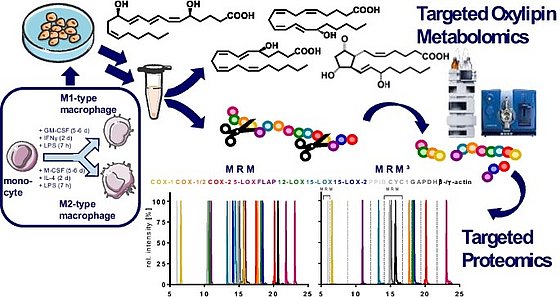
LC-MS/MS quantification is carried out by external calibration using internal standards. A detailed description how to prepare calibration series of the costly and oxidation prone standards and internal standards is described in detail in Koch et al., 2020, excel SI and Hartung et al., 2023. Here it is important to control the actual concentration of standards, because only few certified standards are available. In Hartung et al., 2019 a strategy to determine the concentration of standards. An important parameter for the quality of sample preparation is the recovery of internal standards. For this we suggest the use of internal standards 2 which are described in Koch et al., 2020, excel SI.
Oxylipin formation by oxidative stress
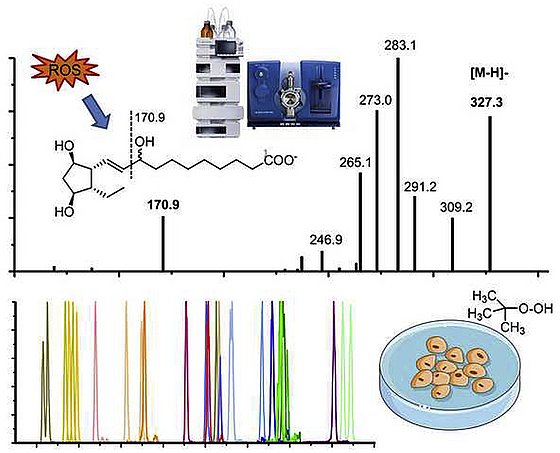
In our method development we set a focus on the analysis of oxylipins formed during oxidative stress. In Rund et al., 2018 the detection of isoprostans and isofurans is described in detail. Moreover Epoxy-PUFA and Dihydroxy-FA are an important new marker of oxidative stress and their detection is described in Rund et al., 2019 and Scholz. et al. 2025, respectively. An SOP for the analysis of both is provided in Rund and Schebb, 2023.
Lipoxins, Resolvins and other so called specialized pro-resolving lipid mediators (SPM)
A large number of studies suggest that specific multiple hydroxylated-PUFA derived from ARA, EPA and DHA play a key role in the resolution of inflammation. Therefore, we developed a method optimized for SPM described. Applying this method, we found that particularly trihydroxylated SPM can not be detected in biological samples, which is described in Kutzner et al., 2019. We also provided a mechanistical explanation for this, showing the formation routes are ineffective in cell free assays (Kutzner et al., 2020), as well as in macrophage (Ebert et al., 2020) and neutrophils (Mainka et al., 2022). This is summarized for leucocytes in Kahnt et al., 2023 and the formation, receptor signaling, formation and detection is questioned in Schebb et al., 2022.
Chiral Analysis
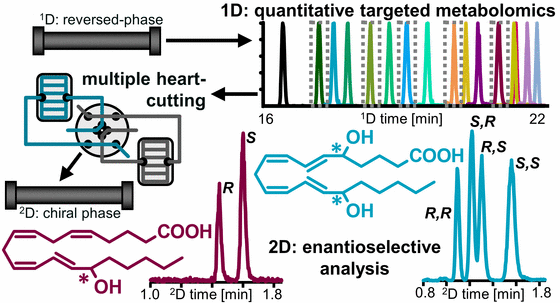
Enzymatic formation of oxylipins occurs stereospecificly, while autoxidation is stereo random. We developed a 2D-LC-MS/MS method for the sensitive enantioselective analysis of hydroxy-PUFA (Kampschulte, 2024). Using that method we could show that 7,17 DiHDHA and 5,15 DiHDHA occurring in human blood are formed during sample preparation and storage and are not the so called RvD5 and RvE4, respectively.
Blood sampling for oxylipin analysis
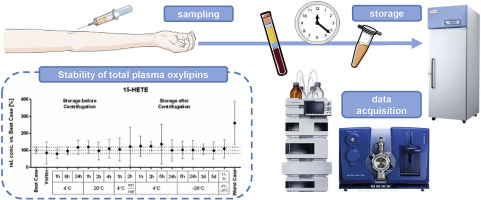
Though oxylipins are paracrine and autocrine acting mediators, the most commonly biological samples analyzed for oxylipins are plasma and serum. For these samples the sampling and pre-analytical sample preparation can influences the measured oxylipin pattern. Oxylipins can degrade and be artificially formed.
The effects of clinical blood sampling, time and procedure of plasma generation on free oxylipins is shown in Rund et al., 2020. The effects of plasma generation and storage on total oxylipins is reported by Koch et al., 2020, excel SI. The expected analytical variability and the intra- and interindividual variability of the plasma oxylipin pattern is described by Ostermann et al., 2018. Based on these information advice for the planning of clinical studies aiming to investigate the oxylipin pattern is given in O´Donnell et al., 2022.
Cell culture experiments
For mechanistical investigations of oxylipin formation cell culture is an indispensable tool. It is crucial to select bovine serum that is suitable for the investigation of oxylipin formation as described by Koch et al., 2021. A strategy to monitor oxylipin formation in the supernatant is shown in Willenberg et al., 2015 and Hartung et al., 2019. Information how to investigate the free oxylipin pattern in cells is described by Hartung et al., 2023 and a detailed SOP summarizes this for both and free total oxylipins can be found in Rund and Schebb, 2023. For the investigation of the n3-PUFA oxylipins we developed an ex-vivo supplementation strategy for primary human macrophages (Kirchhoff et al. 2025).